Hämatologische Neoplasien
Molecular Pathogenesis of Hematological Neoplasias
collaboration between the Institute of Pathology, the CCC-Mainfranken, the Department of Internal Medicine II, the Clinical Research Unit 216 (CRU 216), the “Sander-Therapieeinheit Multiples Myelom”, the “Deutsche Studiengruppe Multiples Myelom” (DSMM) and the International Cancer Genome Consortium (ICGC)
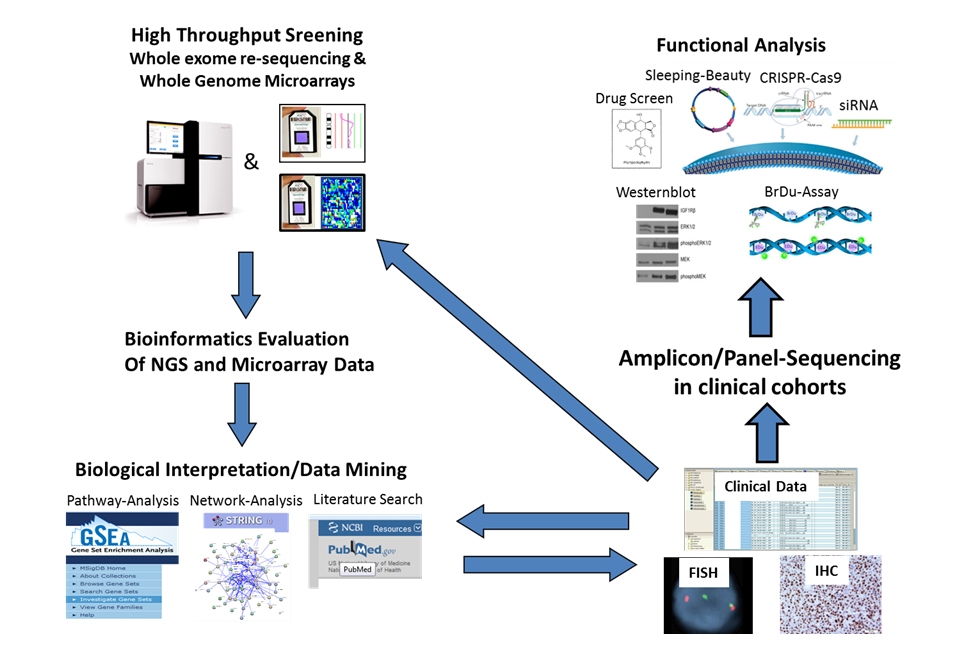
Our group is interested in the molecular pathogenesis of multiple myeloma (MM) and malignant B-cell lymphoma. Specifically, the group focuses on the clinical and molecular characterization of newly detected somatic mutations in MM and the clinical and molecular characterization of t(14;18)-negative follicular lymphoma (FL). Next generation sequencing (NGS) approaches in primary MM and FL patients such as whole exome sequencing and amplicon-sequencing for the detection of single nucleotide variants as well as the integration of gene expression and copy number data are underlying techniques to define and analyze certain molecular subgroups and to identify potential oncogenes that may serve as targets for a more personalized therapeutic approach. Subsequently, in vitro human cell line models are used to better understand the underlying biological processes that are caused by single oncogenic mutations.
Selected Publications
Ellen Leich, Claudia Maier, Riccardo Bomben, Filippo Vit, Alessandro Bosi, Heike Horn, Valter Gattei, German Ott, Andreas Rosenwald, Alberto Zamò Follicular lymphoma subgroups with and without t(14;18) differ in their N-glycosylation pattern and IGHV usage Blood Adv, 2021, ahead of print
Leich E, Schreder M, Pischimarov J, Stühmer T, Steinbrunn T, Rudelius M, Brünnert D, Chatterjee M, Langer C, Keppler S, Heredia-Guerrero SC, Einsele H, Knop S, Bargou RC, Rosenwald A.Novel molecular subgroups within the context of receptor tyrosine kinase and adhesion signalling in multiple myeloma. Blood Cancer J. 2021 Mar 4;11(3):51. doi: 10.1038/s41408-021-00442-2.
Weißbach S, Heredia-Guerrero SC, Barnsteiner S, Großhans L, Bodem J, Starz H, Langer C, Appenzeller S, Knop S, Steinbrunn T, Rost S, Einsele H, Bargou RC, Rosenwald A, Stühmer T, Leich E. Exon-4 Mutations in KRAS Affect MEK/ERK and PI3K/AKT Signaling in Human Multiple Myeloma Cell Lines Cancers (Basel). 2020 Feb 16;12(2):455. doi: 10.3390/cancers12020455. PMID: 32079091
Zamò A, Pischimarov J, Horn H, Ott G, Rosenwald A, Leich E The exomic landscape of t(14;18)-negative diffuse follicular lymphoma with 1p36 deletion Br J Haematol. 2018 Feb;180(3):391-394. doi: 10.1111/bjh.15041.
Zamò A, Pischimarov J, Schlesner M, Rosenstiel P, Bomben R, Horn H, Grieb T, Nedeva T, López C, Haake A, Richter J, Trümper L, Lawerenz C, Klapper W, Möller P, Hummel M, Lenze D, Szczepanowski M, Flossbach L, Schreder M, Gattei V, Ott G, Siebert R, Rosenwald A, Leich E. Differences between BCL2-break positive and negative follicular lymphoma unraveled by whole-exome sequencing. Leukemia. 2018 Mar;32(3):685-693. doi: 10.1038/leu.2017.270. Epub 2017 Aug 21.