Burger Group
RNA Metabolism in Health and Disease
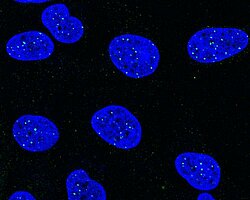
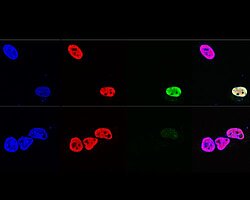
Paraspeckles are phase-separated nuclear bodies in the interchromatin space of mammalian nuclei that are built on two isoforms of the long non-coding RNA NEAT1. More than 40 RNA-binding proteins associate with NEAT1 in a core-shell arrangement. Paraspeckle components serve as multipurpose molecular scaffolds and are involved in various RNA metabolic processes, such as transcriptional regulation, RNA processing, or RNA editing.
NEAT1 serves as a bait for RNA interference factors to promote efficient miRNA biogenesis in unperturbed cells. Replication stress induces the expression of NEAT1 by the tumour suppressor p53, while loss of NEAT1 hyper-sensitises cells to DNA damage. Intriguingly, NEAT1 and other components of paraspeckles are deregulated in cancer and linked to the DNA damage response.
However, the molecular mechanisms that integrate the nuclear RNA metabolism, paraspeckle biology and genome maintenance are poorly understood. We combine basic molecular cell biology and biochemistry with transcriptomics and tumour models to understand the role of paraspeckle components for genome stability and cellular transformation. We aim to decipher the dynamics of paraspeckle composition and discover novel mechanisms that link paraspeckles to genome stability and cellular transformation. We aim to decipher the dynamics of paraspeckle composition and discover novel mechanisms that link paraspeckles to genome stability and tumorigenesis.
We are interested in RNA metabolic events that modulate the stability of the human genome in crosstalk with the DNA damage response. In particular, we aim to understand how the DNA damage response engages RNA metabolic hubs and nuclear bodies in DNA repair. We focus our investigations on RNA-binding proteins and long non-coding transcripts to elucidate RNA-dependent pathways that promote genomic stability in healthy cells and drive tumorigenesis upon deregulation.
The DNA damage response engages components of nuclear paraspeckles to promote genome stability. We aim to understand how RNA-binding proteins respond to oncogenic stress and determine novel functions for RNA-binding proteins in the DNA damage response.
Recent publications
Mamontova V, Trifault B, Boten L, Burger K (2021). Commuting to Work: Nucleolar Long Non-Coding RNA Control Ribosome Biogenesis from Near and Far. Noncoding RNA 7(3):42. [Review] https://doi.org/10.3390/ncrna7030042
Burger K, Schlackow M, Gullerova M (2019). Tyrosine kinase c-Abl couples RNA polymerase II transcription to DNA double-strand breaks. Nucleic Acids Res 47(7):3467-3484. https://doi.org/10.1093/nar/gkz024
Burger K, Gullerova M (2018). Nuclear re-localization of Dicer in primary mouse embryonic fibroblast nuclei following DNA damage.PLoS Genet 14(2):e1007151. https://doi.org/10.1371/journal.pgen.1007151
Burger K, Schlackow M, Potts M, Hester S, Mohammed S, Gullerova M (2017). Nuclear phosphorylated Dicer processes double-stranded RNA in response to DNA damage. J Cell Biol 216(8):2373-2389. https://doi.org/10.1083/jcb.201612131
Kaspar Burger

Junior Group Leader
I studied biology at the Ludwig Maximilian University of Munich, Germany (2003-2009). In 2013, I completed my PhD thesis working on Cdk9 and human ribosome biogenesis in Dirk Eick’s lab and held a fellowship from the German José Carreras Leukaemia Foundation from 2009 to 2011. For my Postdoc, I joined Monika Gullerova’s lab at the William Dunn School of Pathology, University of Oxford, UK to study nuclear Dicer functions in mammals (2013- 2019).
In 2019, I accepted a position as junior group leader at the MSNZ in Würzburg, where we are interested in the regulatory principles that link the RNA metabolism with genome stability and tumour immunology. I investigate the production and processing of RNA in response to DNA damage, with focus on transcription, nuclear bodies, and RNA-binding proteins.
I was born in Bad Tölz, Bavaria in 1983 and enjoy travelling, cycling, hiking, live music, and art.
Barbara Trifault

PhD Student
In 2009, I have started my scientific education with a Laboratory sciences Baccalaureate followed by a Biotechnology course in France and internships to extend my technical skills (2009-2014). Determined to pursue my theoretical scientific education, I rejoined the Life Sciences course at the University of Versailles Saint Quentin en Yvelines, France from 2014- 2017. During my first year of Master (2017-2018), I studied Genetics, Molecular and Cellular Biology in Paris Saclay University pursuing by a specialization in Cellular Biology at Paris Diderot University, France (2018-2019). For my Master thesis, I joined Dr Mark Scott´s lab at the Cochin Institute in France to study cancer-associated SUMOylation mutants of the tumor suppressor PTEN.
In 2020, I accepted a position as PhD student at the MSNZ Würzburg, Germany in order to understand how nuclear paraspeckles components (RNA-binding-proteins) respond to oncogenic stress and determine novel functions for RNA-binding-proteins in the DNA damage response.
I was born in Aubergenville, France in 1995. I am enjoying travelling, hiking, yoga, live concert, collect vinyl and beer tasting.
Victoria Mamontova

PhD Student
I studied biology in St. Petersburg University, Russia (2013‐2019). In my Bachelor I was interested in cytogenetics and studied social stress in mice. My Master thesis was dedicated to the contribution of methyltransferase Set7/9 to tumorigenesis. I also did a short internship in IMB in Mainz, Germany (summer 2018). During the study I took part in different social events in University, especially I enjoyed organizing science contests for kids. Nowadays I’m involved in a Mentor’s program, where I’m helping the current students on their way to graduate.
I started my PhD in here in Würzburg in August 2020. I’m focusing on the lncRNA NEAT1, which is the core component of paraspeckles. The aim of my project is to study the role of NEAT1 in the DNA damage response. I’m also planning to reveal the interactome of NEAT1 and structural changes of paraspeckles in a context of cancer.
I was born in Saratov, Russia. This place is full of beautiful landscapes. That is why I always liked to spend my free time in nature. I’m also fond of drawing, making origamis, doing yoga and backing stuff.
You can find us here:
Biocenter of the University
MSNZ
Am Hubland
97074 Würzburg